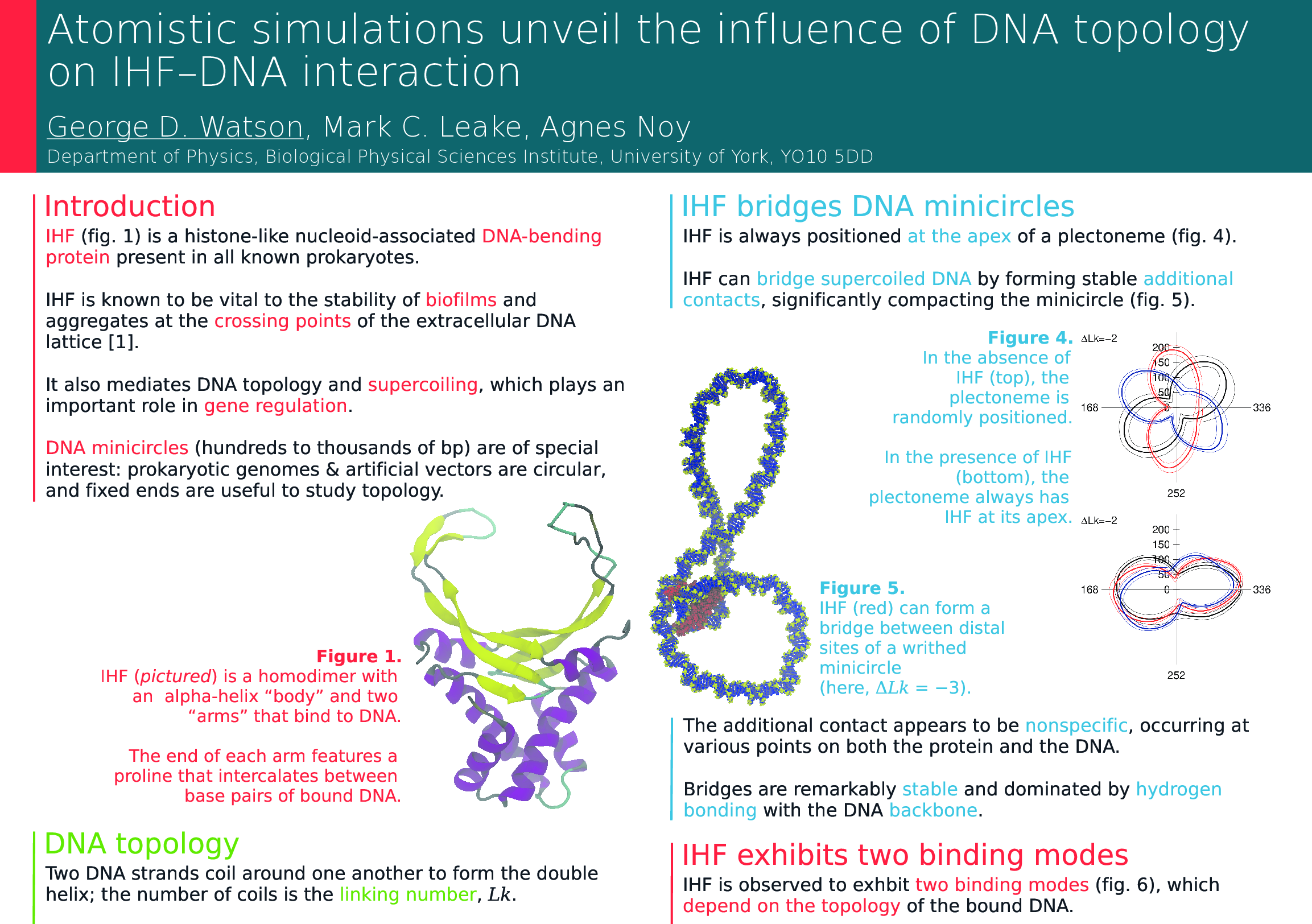
I will be attending the Institute of Physics “Physics of Microorganisms II” conference in London on Monday, with my poster entitled “Atomistic simulations unveil the influence of DNA topology on IHF–DNA interaction”.
Abstract:
IHF is a nucleoid-associated DNA-binding protein that bends DNA by up to 160 degrees and is known to be vital to the stability of bacterial biofilms. Through atomistic molecular dynamics simulations of IHF bound to supercoiled DNA minicircles and linear DNA constructs, the DNA–IHF interaction is studied in unprecedented detail. Novel observations are made of key features of this interaction, including the existence of two clearly distinct binding modes and the formation of bridges between distal DNA sites by IHF, forming closed topological domains. Unlike canonical IHF binding, this bridging appears to be nonspecific, and may be the mechanism by which IHF stabilises crossing points in the extracellular DNA matrix in biofilms. Furthermore, IHF binding both modulates and is modulated by DNA topology, leading to a complex interplay that regulates the bacterial genome and allows regulatory information to be communicated over long distances. Further simulations involving IHF, related proteins, and DNA sequences with multiple binding sites, will soon converge with parallel single-molecule experiments to shed more light on the mechanisms underlying this biologically relevant process and the interactions between nucleoid-associated proteins and DNA.
Watson G D, Leake M C, Noy A
If you’re attending too, find me at board P23.
If you’re not, you don’t need to miss out: A PDF version of my poster is available online.